Medical image analysis
Filter by topics
In collaboration with Centre Hospitalier Universitaire de Lyon, France and University of Patras, Greece.
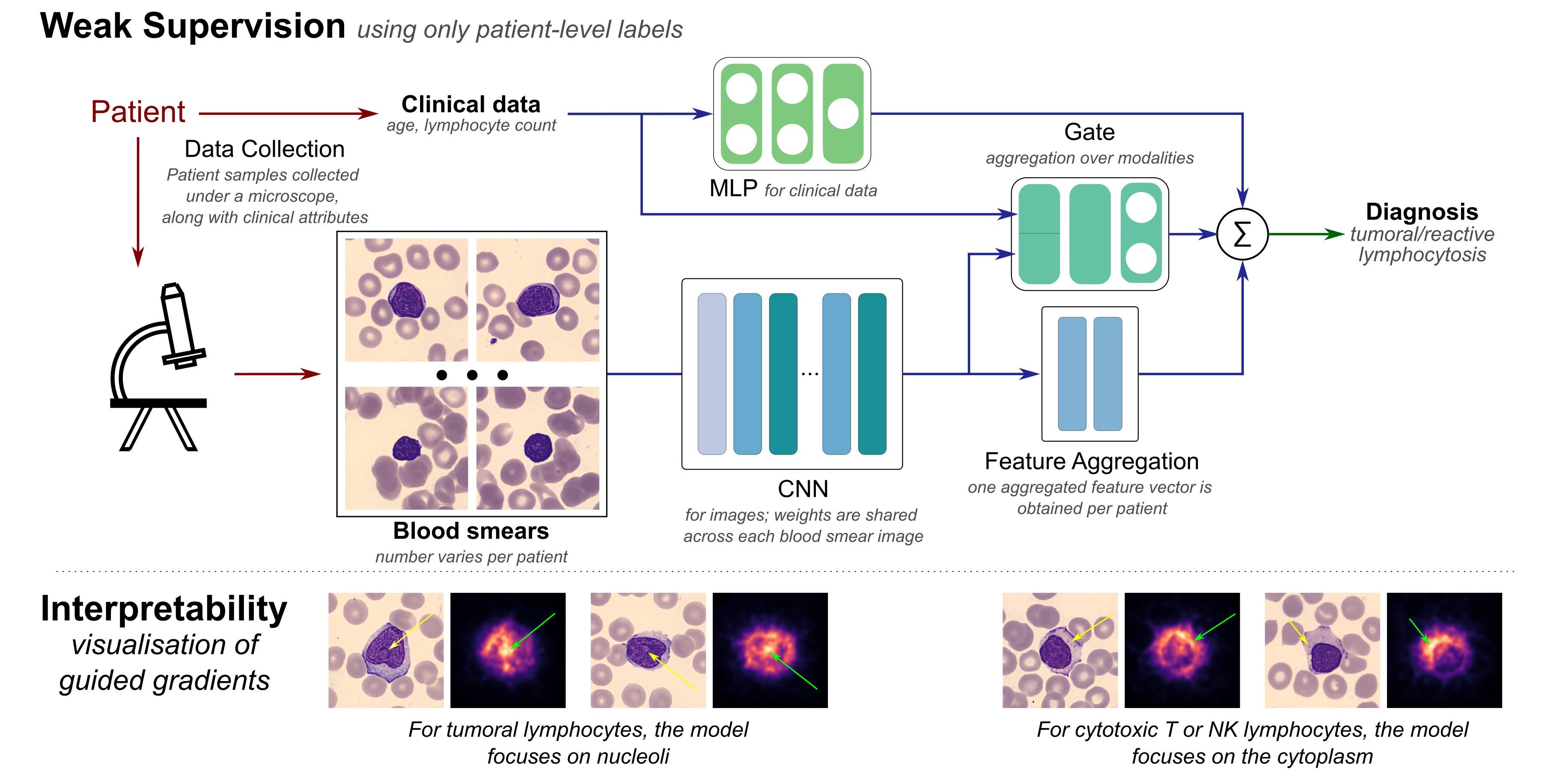
Lymphocytosis (i.e., absolute lymphocyte count above 4 × 109/L) is a common finding in patients, which can be either a reaction to infection, acute stress, and so on (termed reactive), or the manifestation of a lymphoproliferative disorder—a type of cancer of the lymphocytes (termed tumoral). In existing clinical practice, diagnosis (as either reactive or tumoral) relies on visual microscopic examination of blood smears together with the integration of clinical attributes such as age and lymphocyte count. Taking into consideration the visual assessment based on clinical attributes together with texture and size of the lymphocytes in the blood smear, a diagnosis of the subtype of lymphoid malignancy is performed. In this work, we investigate the use of recent advances in deep learning and propose an end-to-end trainable multi-instance convolutional neural network within a mixture-of-experts formulation that combines information from two types of data—blood smears and clinical attributes—for the diagnosis of lymphocytosis. The convolutional network learns to extract meaningful features from images of blood cells using an embedding level approach and aggregates them. Moreover, the mixture-of-experts model combines information from these images as well as clinical attributes to form an end-to-end trainable pipeline for diagnosis of lymphocytosis. Our results demonstrate that even the convolutional network by itself is able to discover meaningful associations between the images and the diagnosis, indicating the presence of important unexploited information in the images. The mixture-of-experts formulation is shown to be more robust while maintaining performance via. a repeatability study to assess the effect of variability in data acquisition on the predictions. Our code and datasets can be found at at this link.
In collaboration with Institut Gustave Roussy, France, Peter MacCallum Cancer Research Centre, Australia, and TheraPanacea, France.
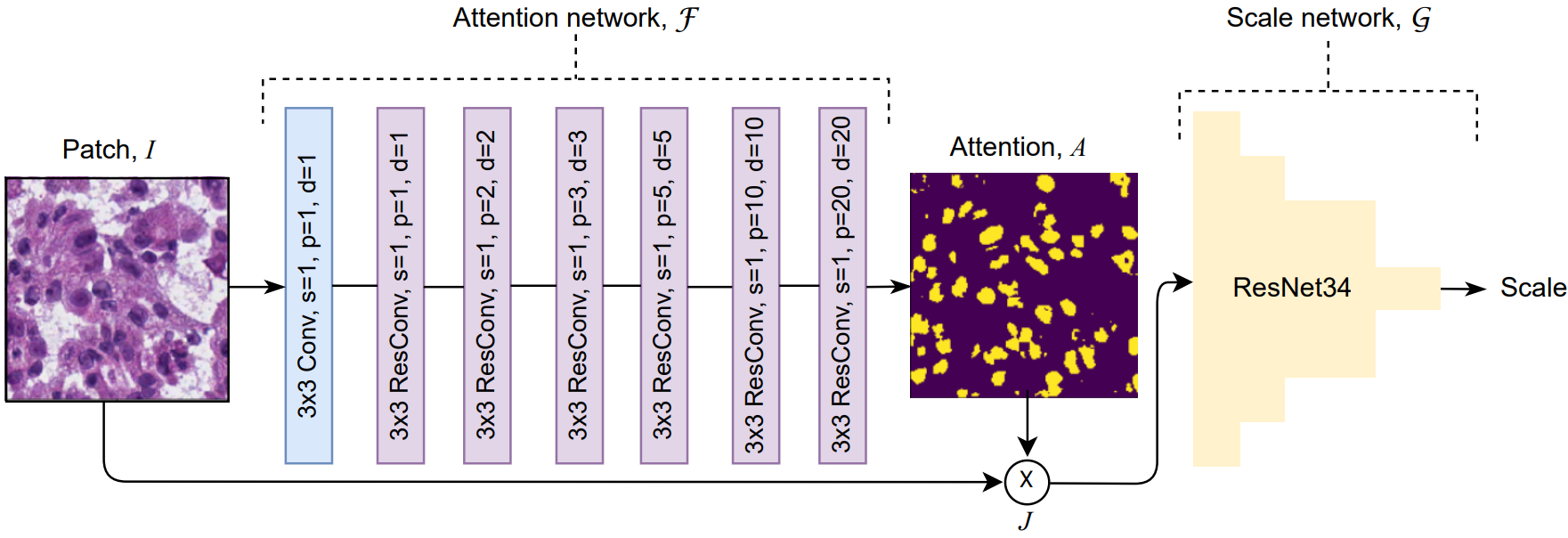
Histology images are the gold standard in diagnosing a considerable number of diseases including almost all types of cancer. For example, the count of nuclei in whole-slide images (WSIs) can have diagnostic significance for numerous cancerous conditions. The proliferation of digital pathology and high-throughput tissue imaging leads to the adoption in clinical practice of digitized histopathological images that are utilized and archived every day. Segmentation and accurate localization of nuclei in histopathological images is a very challenging problem, with most existing approaches adopting a supervised strategy. These methods usually rely on manual annotations that require a lot of time and effort from medical experts. In this study, we present a self-supervised approach for segmentation of nuclei for whole slide histopathology images. Our method works on the assumption that the size and texture of nuclei can determine the magnification at which a patch is extracted. We show that the identification of the magnification level for tiles can generate a preliminary self-supervision signal to locate nuclei. We further show that by appropriately constraining our model it is possible to retrieve meaningful segmentation maps as an auxiliary output to the primary magnification identification task. Our experiments show that with standard post-processing, our method can outperform other unsupervised nuclei segmentation approaches and report similar performance with supervised ones on the publicly available MoNuSeg dataset. Our code and models are available online at this link to facilitate further research.
In this project, we study the impact of deep learning to improve medical imaging registration. We focus on joint training of registration and segmentation network, registration in presence of tumor and clinical application of registration for radiothapy. We apply our algorithms on different cohorts in particular abdomen CT and brain MRI. Our work obtained the 3rd place for Micccai 2020 Learn2Reg Challenge.
In collaboration with the RICE University under the supervision of Prof. Kavraki.
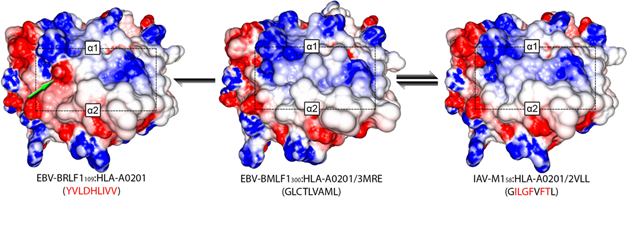
A major challenge in designing immunotherapy is to account for the large variability of possible proteomics profiles of both the particular cancer to treat and of the patient healthy cells. However, most of the methods tackling this task are relying either on the protein sequences that are lacking the ability to consider the spatial structure of the protein or on a biological characterization of the protein structure and then a correlation estimation. We consider HLA proteins for which we have a ground truth dendogram characterizing the similarity between proteins experimentally established. The similarity between the proteins of this dataset has been assessed, in a first step, experimentally by assessing the cross-reactivity to a reference dataset. Graph matching techniques aim to use both information of a graph structure and some features on a graph node to establish a mapping between the nodes of two different graphs. Here, we consider for graphs a mesh on the proteins structures, the nodes being the atoms of the molecule and the edges characterizing spatial proximity of the atoms. The objective of the algorithm is then to maximize the similarity between matched atoms and between the edges of the matched atoms. This objective function is then used to characterize the similarity between the two molecules. Promising results for this proof of concept scenario on the 29 different HLA proteins have been obtained. We achieved an accuracy of 0.83 for the separation of high, low and without cross-reactivity molecules indicating the potentials of our method.
In collaboration with Assistance Publique - Hopitaux de Paris hospitals network.
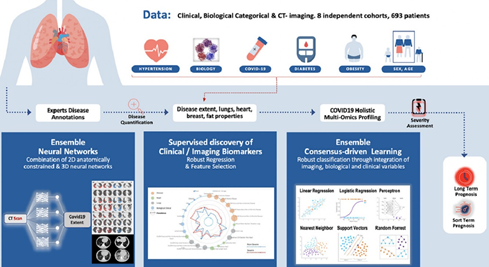
Computed tomography (CT) imaging has been proven to be an important tool for screening, disease quantification and staging. The latter is of extreme importance for organizational anticipation as well as to accelerate drug development through rapid, reproducible and quantified assessment of treatment response. In this study a multi-centric cohort of 536 was considered. We designed an AI driven scheme for the quantification of CT scans for patients suffering from COVID-19 pneumonia. Furthermore, we defined a method for the automatic selection and combination of multi-modal variables towards a holistic signature designed for the COVID-19 triage. On the basis of this interpretable, clinically relevant signature we develop advanced machine learning techniques integrating multi-modal data for severity assessment and short/long term outcome prediction. Our method endows robustness, good generalization properties, explainability and establishes causality with known clinical COVID-19 confounding factors. In conclusion, we show that the combination of chest CT and artificial intelligence can provide tools for fast, accurate and precise disease extent quantification as well as the identification of patients with severe short-term outcomes. Beyond the diagnostic value of CT for COVID-19, our study suggests that AI should be part of the triage process. We are currently working on adapting this pipeline to several other fields such as cancer patients’ response to treatment.
Guillaume Chassagnon, Maria Vakalopoulou, Enzo Battistellaet al., AI-driven quantification, staging and outcome prediction of COVID-19 pneumonia, Medical Image Analysis, Volume 67, 2021

